
H SD and NT activities on a long (2.7 kbp) substrate containing a long pre-formed flap (60 nt) at the end of a 30-nt gap. G Processivities of the strand displacement (SD) and NT reactions in the presence of a Polδ trap competitor: 100 nM Polδ, 100 nM FEN1, 20 ng/µL heparin, and DNA Sub#2 for 30 s at 37 ☌. The NT processing time per nucleotide was calculated as the inverse of the average of the absolute values of the median primer increase and block reduction rates. The experimental datapoints were fitted to linear dependencies with fixed intercepts (28 nt for block length reduction and 33 nt for primer length increase). Median product lengths beyond the first NP were determined for each reaction time as described in “Methods”. F Quantification of the reaction rates presented in panels D and E. D, E NT products monitored through the extension of the primer oligonucleotide ( D) or through the cleavage of the block oligonucleotide ( E): 250 nM Polδ, 250 nM FEN1, RT, DNA Sub#2, and Sub#3 (Supplementary Fig. The experimental datapoints were fitted to a single-exponential product-formation burst equation \([\) (155.2 s) and then the result was divided by 12 nt (the length of the RNA region of the block). C Quantification of the time dependence of the ligated MOF product yield from panel B. B Reaction products for the reconstituted human MOF: 250 nM Polδ, 250 nM FEN1, 250 nM Lig1, and DNA Sub#1 (Supplementary Fig. 1A) 5, 11, 16, 17.Ī Cartoon depiction of the maturation of Okazaki fragments (MOF) and nick translation (NT) reactions. Flap endonuclease-1 (FEN1) cleaves the 5′flap 13, 14, 15 and generates a nick product (NP) that can then be sealed by DNA Ligase 1 (Lig1) (Fig. PCNA encounters the RNA primer on the preceding OF, performing limited strand displacement (SD) synthesis and giving rise to a single-stranded 5′flap structure 11, 12 (Fig.Maturation of Okazaki fragments (MOF) is initiated as Polδ Thereafter, RFC loads PCNA onto the primer-template (P/T) junction 10 for enhanced Polymerase δ (Polδ) processivity during primer extension. 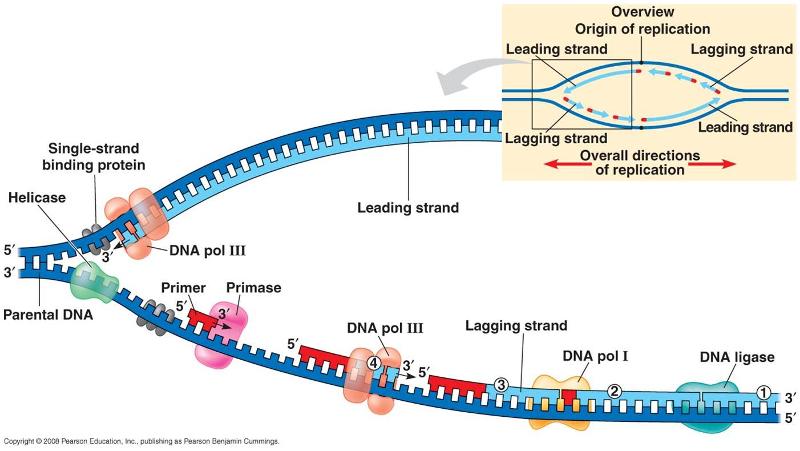
OF synthesis is initiated by the polymerase α (Polα)-primase complex, which generates a hybrid primer of 8–12 RNA and 10–20 DNA nucleotides 8, 9. The leading strand is continuously replicated, while lagging strand synthesis is discontinuous, via the formation of short Okazaki fragments (OFs), extending for ~200 nucleotides (nt) 7. The unidirectional synthesis by DNA polymerases and the chemical bidirectionality of DNA force the replisome to copy parental strands via two distinct modes 1, 2, 3, 4, 5, 6.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
